Plant Computational Biology
Plants have evolved organs, specialized metabolites, and a myriad of physiological and molecular adaptive strategies to thrive in adverse environments. A better understanding of the genes and pathways behind these adaptive strategies would provide unprecedented knowledge of how plants evolve and how to engineer more stress-resilient crops with higher nutritional values. However, the function of most genes in the plant kingdom is unknown, constrained to only a handful of model organisms, or inferred by genomic approaches, which often cannot reveal which genes work together in a biological pathway.
 Our long-term goal is to uncover the function and evolution of genes, stress resilience mechanisms, and specialized metabolic biosynthetic pathways of algae and land plants by a combination of comparative experimental and computational approaches.
Our long-term goal is to uncover the function and evolution of genes, stress resilience mechanisms, and specialized metabolic biosynthetic pathways of algae and land plants by a combination of comparative experimental and computational approaches.
For up to date group news, please visit https://publish.obsidian.md/mutwillab/Homepage/News
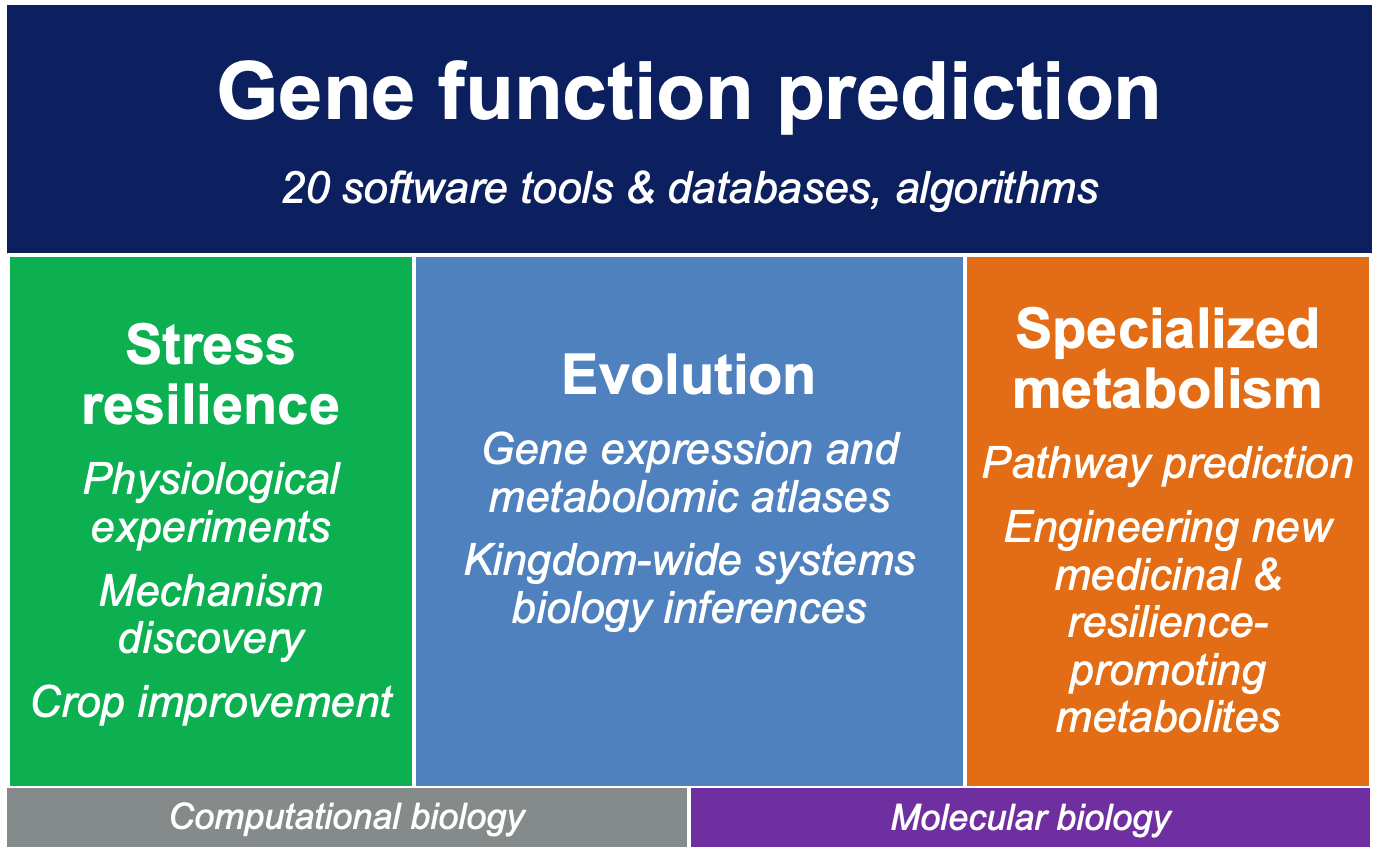
Our group employs four interconnected and cross-disciplinary research pillars, combining experimental and computational approaches:
- Evolution: Investigate how multi-omics approaches can reveal the evolution of genes, pathways, and traits across the plant kingdom.
- Stress: Explore plant responses to stress combinations, organ-specific dynamics, and multi-layered gene regulation in diverse species.
- Specialized Metabolism: Identify and engineer specialized metabolic pathways using genomic and chemoinformatic methods.
- Tools & Databases: Develop advanced computational tools and online databases to provide the plant community with robust gene function predictions and resources.
The methods used in the group include:
- Genome and transcriptome sequencing, assembly and anntotation
- Construction and analysis of biological networks
- Physiological experiments on plants and sampling of plant material from field
- Usage and development of biostatistical models, machine learning and deep learning approaches, artificial intelligence, including large language models
- Generation of transgenic plants, bacteria, fungi and mutant analyses
We have produced several high impact publications associated with the four research pillars.
- Evolution: a database to study gene expression in the plant kingdom (Koh et al., 2024, TPJ), cover of ancestral genome reconstruction paper (Mutwil, Fernie, 2024, MP), review on kingdom-wide expression analyses (Julca et al., 2023, TIPS), analysis of the evolution of plant organs (Julca et al., 2021, NPlants), and the evolution of ferns (Ali et al., 2025, NPlants)
- Abiotic stress responses: review on gene expression data and AI (Koh et al., 2024, CSBJ), responses to combined stress (Tan et al., 2023, NComm),
- Specialized metabolism: review on sesquiterpene lactones (Agatha et al., 2024, PTRSLBBC), antibacterial activites in Melastoma (Poh et al., 2024, FIPS), review on chemoinformatics (Lim et al., 2024, CSBJ), anticancer activity of Oldenlandia (Julca et al., 2023, JIPB), review on gene expression data usage (Delli-Ponti et al., 2021, FIPS),
- Bioinformatical tools: review on large language models (Sunil et al., 2024, COPB), diurnal gene expression database (Tan et al., 2024, PCP), de novo transcriptome assembly (Lim et al., 2024, PPlantarum), review on large language models (Lam et al., 2024, TIPS), correlation of biological features (Ng et al., 2022, FIPS), review on plant trascriptomics (Lim et al., 2022, PComm), database for protists (Villanueva et al., 2022, JBM) and bacteria (Lim et al., 2022, JMB), structure of virus genomes (Delli-Ponti, Mutwil, 2021, BMC G), assembly of gene expression data in plant kingdom (Goh, Mutwil, 2021, Bioinformatics),
Group members
| Name | Title | Phone | |
|---|---|---|---|
| Search in Name | Search in Title | Search in Phone | |
| Jia Min Lee | Postdoc | +4535333440 | |
| Mads Harder Møller | PhD Fellow | +4535335179 | |
| Marek Mutwil | Associate Professor | +4535326080 |
External members
Mutwil is also responsible for group members in Singapore.
| Name | Titel |
|---|---|
| Dr. Eugene Koh | Postdoc |
| Ms. Olivia Agatha | PhD student |
| Mr. Peng Ken Lim | PhD student |
| Mr. Shan Chum Lim | PhD student |
| Ms. Jooa Moon | PhD student |
| Mr. Manoj Itharajula | PhD student |

Research group leader
Marek Mutwil
Associate Professor
mutwil@plen.ku.dk
+45 35 32 60 80
